GARVAN INSTITUTE OF MEDICAL RESEARCH—Findings from the Garvan Institute of Medical Research show modern humans acquired a gene variant from Denisovans that heightened their immune reactions, indicating adaptation of the immune system to a changing environment.
The breakthrough study, published in Nature Immunology, is the first to demonstrate a single DNA sequence variant from an extinct human species that changes the activity of the modern human immune system.
The Denisovans – an extinct human species related to Neanderthals – interbred with modern humans ~50,000 years ago during the migrations of modern humans from Africa to what is now Papua New Guinea and Australia. Today, up to 5% of the genome of people indigenous to Papua New Guinea is derived from Denisovans.
The Garvan study reveals that modern humans acquired a gene variant from Denisovans that increases a range of immune reactions and inflammatory responses – including reactions that protect humans from disease-causing microbes.
“Our study indicates that the Denisovan gene variant heightens the inflammatory response in humans,” says co-senior author Associate Professor Shane Grey, who heads the Transplantation Immunology Laboratory at Garvan.
“Previous research has found collections of gene variants from extinct human species that appear to have provided an advantage to humans living at high altitudes or to resist viruses, but have been unable to pinpoint which if any were actually functional,” he adds. “This study is the first to identify a single, functional variant, and suggests that it also had an evolutionary benefit on the human immune system.”
Discovering an immune switch
Harmful versions of a gene called TNFAIP3 have long been associated with the overactive immunity in autoimmune conditions, including inflammatory bowel diseases, arthritis, multiple sclerosis, lupus, psoriasis and type 1 diabetes. The TNFAIP3 gene codes for a protein called A20 that helps ‘cool’ the immune system by reducing immune reactions to foreign molecules and microbes.
As part of a collaboration between Garvan, the Sydney Children’s Hospital, Randwick, the Children’s Hospital at Westmead, and the Clinical Immunogenomics Research Consortium of Australasia (CIRCA), the researchers analysed the genomes of families in which one child presented with a severe and unusual autoimmune or inflammatory condition.
“Four separate families had the same DNA variant in the TNFAIP3 gene, changing one amino acid in the A20 protein from an isoleucine to a leucine (I207L)”, says Professor Goodnow, Executive Director of the Garvan Institute and co-senior author of the study. “However, the presence of this variant in healthy family members indicated it was not sufficient to cause inflammatory disease on its own.”
The researchers extracted immune cells from the families’ blood samples, and found that, in cell culture, they produced a stronger inflammatory response than the immune cells of other individuals.
Tracing back immunity
Using datasets made available through the Simons Genome Diversity Project, the Indonesian Genome Diversity Project, Massey University, and the Telethon Kids Institute, which includes genome sequence data on hundreds of diverse human populations, co-first author and Flinders University senior researcher Dr Owen Siggs investigated the worldwide distribution of the TNFAIP3 variant.
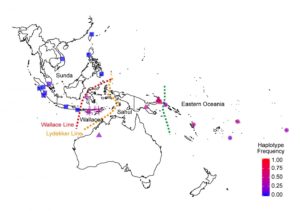
The I207L variant carried by the Sydney families was absent from most populations but common in indigenous populations east of the Wallace Line, a deep ocean trench passing between Bali and Lombok and separating Asian fauna to the west from Australian fauna to the east. The I207L variant was common in people throughout Oceania, including people with Indigenous Australian, Melanesian, Maori and Polynesian ancestry.
“The fact that this rare version of the gene was enriched in these populations, and displayed genetic signatures of positive selection, means it was almost certainly beneficial for human health,” says Associate Professor Grey.
The team also discovered the I207L variant in the genome sequence of an extinct human species, extracted from a 50,000-year-old finger bone of a Denisovan girl, found inside the Denisova cave in the Altai Mountains of Siberia. “Making that connection was extremely exciting,” says Dr Siggs.
The I207L variant was present in two copies in the Denisovan girl but absent from Neanderthal remains from the same cave, indicating that the immunity-enhancing gene variant arose after the divergence of the Denisovan and Neanderthal lineages ~400,000 years ago.
Dialling up the immune system
To investigate the Denisovan gene variant’s effects on the immune system, co-first author Dr Nathan Zammit replicated the I207L variant in a mouse model. “When exposed to a pathogenic Coxsackie virus strain – a virus which was originally isolated from a fatal case of human infant infection – mice with the Denisovan variant had stronger immune reactions and resisted the infection better than mice without the Denisovan gene,” Dr Zammit explains.
“Our study indicates that the Denisovan variant, and others like it, act on a ‘temperature control’ dial in the immune system, turning up the temperature to change how we respond to different microbes,” says Professor Goodnow.
“It was previously thought that A20, a gene that’s central to the immune system, is binary – either it’s switched on or off,” adds Associate Professor Grey. “We’ve found it in fact tunes us as individuals to optimal ‘Goldilocks points’ in between – where immune reactions are neither too hot nor too cold – and that blows the field wide open.”
____________________________

Map shows the frequency of the Denisovan TNFAIP3 gene variant in modern human populations of Island South East Asia and Oceania, it is found to be common east of the Wallace Line. Owen Siggs
____________________________
Article Source: GARVAN INSTITUTE OF MEDICAL RESEARCH news release





