UNIVERSITY OF UTAH—Aug. 7, 2017—Hundreds of thousands of years ago, the ancestors of modern humans diverged from an archaic lineage that gave rise to Neanderthals and Denisovans. Yet the evolutionary relationships between these groups remain unclear.
A University of Utah-led team developed a new method for analyzing DNA sequence data to reconstruct the early history of the archaic human populations. They revealed an evolutionary story that contradicts conventional wisdom about modern humans, Neanderthals and Denisovans.
The study found that the Neanderthal-Denisovan lineage nearly went extinct after separating from modern humans. Just 300 generations later, Neanderthals and Denisovans diverged from each other around 744,000 years ago. Then, the global Neanderthal population grew to tens of thousands of individuals living in fragmented, isolated populations scattered across Eurasia.
“This hypothesis is against conventional wisdom, but it makes more sense than the conventional wisdom.” said Alan Rogers, professor in the Department of Anthropology and lead author of the study that will publish online on August 7, 2017 in the Proceedings of the National Academy of Sciences.
A different evolutionary story
With only limited samples of fossil fragments, anthropologists assemble the history of human evolution using genetics and statistics.
Previous estimates of the Neanderthal population size are very small — around 1,000 individuals. However, a 2015 study showed that these estimates underrepresent the number of individuals if the Neanderthal population was subdivided into isolated, regional groups. The Utah team suggests that this explains the discrepancy between previous estimates and their own much larger estimate of Neanderthal population size.
“Looking at the data that shows how related everything was, the model was not predicting the gene patterns that we were seeing,” said Ryan Bohlender, post-doctoral fellow at the M. D. Anderson Cancer Center at the University of Texas, and co-author of the study. “We needed a different model and, therefore, a different evolutionary story.”
The team developed an improved statistical method, called legofit, that accounts for multiple populations in the gene pool. They estimated the percentage of Neanderthal genes flowing into modern Eurasian populations, the date at which archaic populations diverged from each other, and their population sizes.
A family history in DNA
The human genome has about 3.5 billion nucleotide sites. Over time, genes at certain sites can mutate. If a parent passes down that mutation to their kids, who pass it to their kids, and so on, that mutation acts as a family seal stamped onto the DNA.
Scientists use these mutations to piece together evolutionary history hundreds of thousands of years in the past. By searching for shared gene mutations along the nucleotide sites of various human populations, scientists can estimate when groups diverged, and the sizes of populations contributing to the gene pool.
“You’re trying to find a fingerprint of these ancient humans in other populations. It’s a small percentage of the genome, but it’s there,” said Rogers.
They compared the genomes of four human populations: Modern Eurasians, modern Africans, Neanderthals and Denisovans. The modern samples came from Phase I of the 1000-Genomes project and the archaic samples came from the Max Planck Institute for Evolutionary Anthropology. The Utah team analyzed a few million nucleotide sites that shared a gene mutation in two or three human groups, and established 10 distinct nucleotide site patterns.
Against conventional wisdom
The new method confirmed previous estimates that modern Eurasians share about 2 percent of Neanderthal DNA. However, other findings questioned established theories.
Their analysis revealed that 20 percent of nucleotide sites exhibited a mutation only shared by Neanderthals and Denisovans, a genetic timestamp marking the time before the archaic groups diverged. The team calculated that Neanderthals and Denisovans separated about 744,000 years ago, much earlier than any other estimation of the split.
“If Neanderthals and Denisovans had separated later, then there ought to be more sites at which the mutation is present in the two archaic samples, but is absent from modern samples,” said Rogers.
The analysis also questioned whether the Neanderthal population had only 1,000 individuals. There is some evidence for this; Neanderthal DNA contains mutations that usually occur in small populations with little genetic diversity.
However, Neanderthal remains found in various locations are genetically different from each other. This supports the study’s finding that regional Neanderthals were likely small bands of individuals, which explains the harmful mutations, while the global population was quite large.
“The idea is that there are these small, geographically isolated populations, like islands, that sometimes interact, but it’s a pain to move from island to island. So, they tend to stay with their own populations,” said Bohlender.
Their analysis revealed that the Neanderthals grew to tens of thousands of individuals living in fragmented, isolated populations.
“There’s a rich Neanderthal fossil record. There are lots of Neanderthal sites,” said Rogers. “It’s hard to imagine that there would be so many of them if there were only 1,000 individuals in the whole world.”
Rogers is excited to apply the new method in other contexts.
“To some degree, this is a proof of concept that the method can work. That’s exciting,” said Rogers. “We have remarkable ability to estimate things with high precision, much farther back in the past than anyone has realized.”
____________________________________
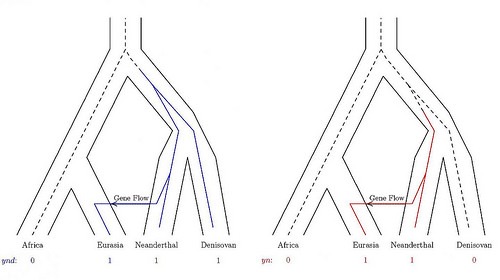
These population trees with embedded gene trees show how mutations can generate nucleotide site patterns. The four branch tips of each gene tree represent genetic samples from four populations: modern Africans, modern Eurasians, Neanderthals, and Denisovans. In the left tree, the mutation (shown in blue) is shared by the Eurasian, Neanderthal and Denisovan genomes. In the right tree, the mutation (shown in red) is shared by the Eurasian and Neanderthal genomes. Credit: Alan Rogers, University of Utah
_________________________________________________
Article Source: University of Utah news release
_________________________________________________
Chad Huff of the Department of Epidemiology at the MD Anderson Cancer Center at the University of Texas also contributed to the study. This work was supported by the National Science Foundation (BCS 1638840), the Center for High Performance Computing at the University of Utah and the National Cancer Institute (R25CA057730 and CA016672).
_________________________________________________
Receive 30 days free access to the popular new CuriosityStream lineup of documentaries on science, history, nature, and technology as a new Popular Archaeology premium subscriber.
___________________________________________
Travel and learn with Far Horizons.
____________________________________________
This richly illustrated issue includes the following stories: Recent findings shedding new light on the whereabouts of the remains of Philip of Macedon, father of Alexander the Great; how an archaeologist-sculptor is bringing bones of the dead back to life; archaeologists uncovering town life at the dawn of civilization; an exclusive interview with internationally acclaimed archaeologist James M. Adovasio about what makes the Meadowcroft Rockshelter prominent in the ongoing search for the first Americans; what archaeologists are finding at the site of the ancient city of Gath, the home town of the biblical Philistine giant, Goliath; and how scientists are redrawing the picture of human evolution in Europe. Find it on Amazon.com.