Standing on stage, Svante Pääbo makes a soft-spoken yet commanding impression before his audience. Dressed in a loose, well-pressed suit with eyeglasses that make him appear the stereotype of the quintessential scientist, he expounds on a topic that has recently commanded headlines and captured imaginations the world over—how the field of genetics has been illuminating and even revolutionizing what we know about human evolution and the human prehistoric past.
Pääbo is a director at the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany. He and his team are known for developing a methodology for isolating and sequencing actual ancient DNA by sampling the remains of extinct human species. He is perhaps best known for his genetic findings suggesting that Neanderthals, an extinct form of human that lived in Eurasia more than 30,000 years ago, mated with modern humans and left their genetic imprint in people who live today.
“Neanderthals are not totally extinct,” he said before his audience. “In some of us they live on, a little bit.”
Pääbo was speaking in this instance before an audience at the international TED (Technology, Entertainment, Design) conference held in Edinburgh, Scotland in July, 2011. Entitled “DNA Clues to Our Inner Neanderthal”, he explained how he and his team of scientists have been able to look into the complex elements of the human genome and ferret out the telltale signs of these prehistoric human cousins within our own ancestry.
But Pääbo spoke of much more than Neanderthals. “From a genomic perspective, we are all Africans,” he said with clear assurance—his work has also helped to show that all humans can trace their ancestry far back to a small population of Africans who later expanded and spread out across the globe. It has become a widely accepted paradigm among anthropologists and geneticists worldwide. Being at the forefront of that paradigm, he has, arguably more than any other scholar, become in some ways an iconic figure of the field of paleogenetics, the study of the past through the analysis of preserved genetic material from the remains of ancient organisms, a field that has become somewhat romanticized and sensationalized through popular cinematic productions like Jurassic Park
_______________________________________________
But it didn’t all begin with Pääbo. Emile Zuckerkandl and the physical chemist Linus Carl Pauling introduced the term “paleogenetics” in 1963, and in 1984 a team led by Allan Wilson published the achievement of the first sequence of ancient DNA isolated from the remains of the quagga, an extinct form of zebra.[1] Wilson’s 1984 breakthrough kick-started a scientific explosion of efforts and studies that would take the field to new technical breakthroughs, using developing new laboratory techniques, methods of DNA abstraction and complex computational tools.
Finding Ancient Code in the Living
Recovering and studying ancient DNA (aDNA) from long-dead organisms tells only part of the story. There is a history buried deep within our living genes, a history that could possibly take us back to glimpse the primordial soup of our human ancestry, perhaps even as far back as the ancient proto-humans who walked the world long before modern humans. It can be found in the “fossil” shadows of our past that actually exist today within our genes—clues that have fascinating implications for human evolution and past population migrations and dispersals.
Knowing that many current, ‘living’ genomic sequences existing in today’s populations are also replicated copies of DNA that existed in the past, researchers over the past few decades began using concepts and models such as ‘coalescent theory’ and ‘multilocus nested clade analysis’ (MLNCA) on the mitochondrial genome (mtDNA) and the Y chromosome, those parts of the genome that have seen little or no recombination (mixing with other DNA segments that have a different evolutionary history). It was much like studying “fossil” DNA imprinted within the genome. In this way, scientists were able to discover what is famously penned today in the popular culture as “Mitochondrial Eve”, the matrilineal most recent common ancestor (MRCA), which suggested that modern humans descended in a direct, unbroken, maternal line to a single female ancestor who lived approximately 100,000–200,000 years ago, most likely in East Africa. The discovery was made in the 1980’s thanks to Allan Wilson of the University of California, Berkeley, along with doctoral students Mark Stoneking and Rebecca L. Cann, who developed and used a molecular clock, a technique by which species divergence time could be estimated.
In other words, we were all originally Africans. It became a headline-making story.
But there was more. Alan R. Templeton, the Charles Rebstock Professor of Biology and Genetics and Professor of Evolutionary and Environmental Biology in the Department of Biology at Washington University in St. Louis, in an article[1] published in a 2013 issue of Genetic Engineering and Biotechnology News, has summarized some of the suggestions that have emerged—human population and evolution ‘events’ through time that have defined the directions of our ancestry. He points to three major ‘events’ that have been suggested by genetic research.
“The first detected event,” he writes, “ is an expansion out of Africa into Eurasia dated to 1.9 million years ago using a molecular clock—an inference confirmed by the fossil record.”* The landmark fossil discoveries of 1.8 million-year-old hominins in Dmanisi, Georgia, by David Lordkipanidze of the Georgian National Museum, are prime examples supporting this finding. A second dispersal out of Africa event occurred about 650,000 years ago, also attested to by the archaeological record—such as the Acheulean stone tool industry with its most characteristic marker, the bifacial hand-axe, and the fossil record of hominins such as Homo erectus, traditionally thought to be the first global colonizing human ancestor. Finally, Templeton suggests that a third dispersal out of Africa occurred about 130,000 to 125,000 years ago, reaching East Asia by about 110,000 years ago.
This third major dispersal has also been perhaps the most debated. Some recent studies, however, have produced results that may lend further support to the theory. One such study involved an international team of scientists conducting an analysis of the genetic diversity and cranial measurements of 10 African and Asian human populations, concluding that anatomically modern (AMH) humans may have dispersed out of Africa earlier than previously thought, and in more than one stage: initially into Asia by taking a southern route through Arabia as much as 130,000 years ago; and later into Northern Eurasia on a more northerly route 50,000 years ago. As the timing and nature of early modern human dispersal out of Africa has long been disputed among scholars, with competing theories or models about how and when it all occurred, the research team analyzed their data within the framework of the competing models. They came up with the model that best fits the results. “We tested for the first time to our knowledge the spatiotemporal dimensions of competing out-of-Africa dispersal models,” write Hugo Reyes-Centeno and colleagues in their report, “analyzing in parallel genomic and craniometric data. Our results support an initial dispersal into Asia by a southern route beginning as early as ∼130 ka and a later dispersal into northern Eurasia by ∼50 ka.”* Reyes-Centeno is a paleoanthropologist with the Senckenberg Center for Human Evolution and Paleoenvironment, Eberhard Karls University of Tübingen, Germany. Other researchers included Katerina Harvati, also of Eberhard Karls University; Silvia Ghirotto and Guido Barbujani of the University of Ferrara, Ferrara, Italy; and Florent Détroit and Dominique Grimaud-Herve of the National Museum of Natural History, Paris, France. “This is consistent with archaeological evidence for modern human occupation in the southern Arabian Peninsula at ∼125 ka,” wrote the authors. “This date [130,000 ka] is in intriguingly closer correspondence with the genetic divergence estimates for our sampled populations, with a calendar date of divergence between Melanesians and South Africans at ∼116 ka, for example. No modern human fossils have been discovered in the southern Arabian Peninsula, but lithic artifacts show affinities with African assemblages, including those discovered alongside the fossil remains at Herto, Ethiopia, dated between ∼154–160 ka.”**
Extracting Ancient Genes from the Bones
Most scientists would arguably agree, however, that there is nothing quite like getting the direct evidence from the ancient specimens themselves. They knew that extracting actual aDNA from human bones dated to prehistoric periods, in a very real sense, would move us beyond the ‘smoking gun’ of the ‘fossil shadows’ imprinted within our genes. It was therefore a natural step to pick the available human fossil bones themselves of their DNA and to study them.
It has not been an easy task. The challenge has been fraught with technical, methodological, and environmental difficulties and complexity, which has included the tricky business of dealing with contamination from other DNA introduced into the sample study mixes as a result of such things as modern human handling and microbes, as well as the fact that ancient DNA has had tens of thousands of years to break down or ‘deteriorate’ within the environmental context since death of the individuals. Nevertheless, geneticists and other specialists have continued to advance the science with improved techniques, technologies, and knowledge.
The Neanderthals
No ancient hominin fossils have been subject to as much DNA sampling and analysis than that of our famous ancient human cousins, the Neanderthals. Through the study of bone samples from a number of Neanderthal specimens recovered from different archaeological sites across Eurasia, a vast dataset has been developed, characterizing the Neanderthal genome as the best known and most complete archaic human genome of any extinct hominin. Genetically, we only know more about one other hominin—modern humans, the last and only surviving hominin on the planet. Pivotal to the Neanderthal research was a landmark study published in Science in May of 2010, where R.E. Green, Svante Pääbo and colleagues at the Max-Planck Institute for Evolutionary Anthropology in Leipzig, Germany, relate in detail how they generated a draft sequence of the Neanderthal genome composed of more than 4 billion nucleotides from three Neanderthal bone specimens. [2] They compared the Neanderthal genome to the genomes of five present-day humans from different parts of the world, identifying genomic regions that were possibly affected by selection in ancestral modern humans, genes such as those involved in metabolism and cognitive and skeletal development. In the process, they were able to show that Neanderthals shared more genetic variants with present-day humans in Eurasia than with present-day humans in sub-Saharan Africa, suggesting that gene flow from Neanderthals to the ancestors of present-day non-Africans likely occurred before the divergence of Eurasian groups from each other.
___________________________________________
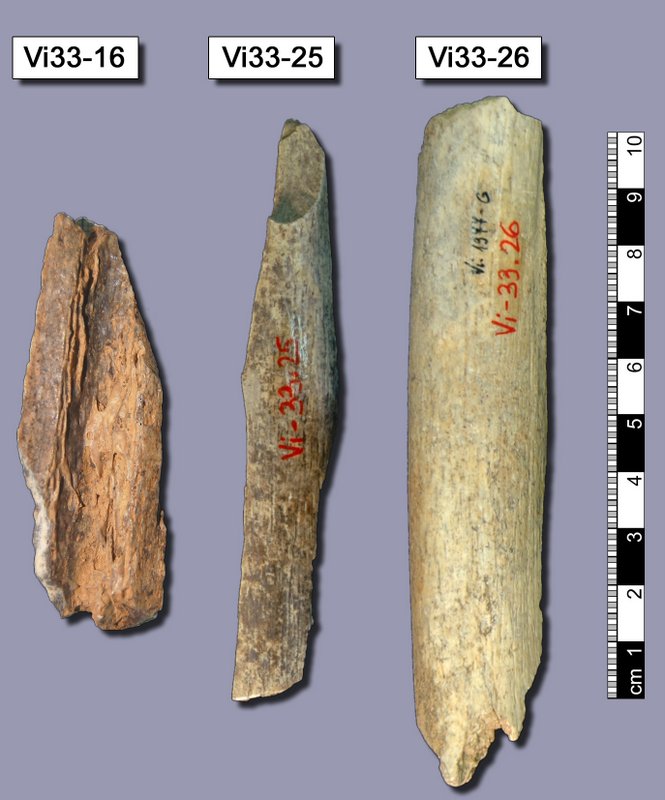 Three bones from Vindija cave, Croatia. Most of the Neanderthal genome sequence was retrieved from these bones. This image relates to the paper published in Science on 7 May 2010 by Richard Green and colleagues titled “A Draft Sequence of the Neandertal Genome.” This image relates to a paper that appeared in the Dec. 23, 2011, issue of Science, published by AAAS. The paper, by Science News staff in Washington D.C., was titled, “Breakthrough of the Year.” Image courtesy of Max-Planck-Institute EVA
Three bones from Vindija cave, Croatia. Most of the Neanderthal genome sequence was retrieved from these bones. This image relates to the paper published in Science on 7 May 2010 by Richard Green and colleagues titled “A Draft Sequence of the Neandertal Genome.” This image relates to a paper that appeared in the Dec. 23, 2011, issue of Science, published by AAAS. The paper, by Science News staff in Washington D.C., was titled, “Breakthrough of the Year.” Image courtesy of Max-Planck-Institute EVA
_______________________________________________________
As research has progressed, geneticists and other scientists have been able to piece together some interesting suggestions for four evolutionary traits related to this, what is considered modern humanity’s closest extinct hominin relative:
First, sequencing indicated that Neanderthal and modern human populations split from a common ancestor about ca. 550,000 – 765,000 years ago. Second, although Neanderthals are thought to have mostly inhabited what is present-day Eurasia and the Levant, genetic evidence of their presence has also been found as far east as the Okladnikov and Denisova caves in the Altai mountains of Western Siberia, suggesting the existence of distinct groups across a very broad geographic range. Third, even given the broad range, the size of the Neanderthal population seems to have been much smaller than that of the later, emerging modern humans, effectively only one-tenth the size of the latter. And fourth, more than their modern human cousins, the Neanderthals seem to have engaged in more intra-familial mating patterns, such as inbreeding between half-siblings, uncle and niece, grandfather and granddaughter, or double first cousins. Some scholars hypothesize that this may be in part because of the smaller size of the population groups and their comparatively greater extent of isolation as compared with their modern human counterparts.
Certainly a great deal more needs to be researched related to the Neanderthal genome, and ongoing and future studies may indeed challenge this emerging picture. But it wouldn’t be an exaggeration to say that the pace and volume of new findings about the Neanderthals have been nothing less than remarkable.
The Denisovans
In March 2010, scientists announced the discovery of an ancient finger bone fragment from a female who lived in the remote Denisova Cave in the Altai Mountains in Siberia, a cave where archaeological excavations had yielded fossils of Neanderthals and prehistoric modern humans. But this finger bone, along with two teeth and a toe bone that were recovered later, came from a human that was neither Neanderthal nor modern human. Analysis of the mitochondrial DNA (mtDNA) showed this individual to be genetically distinct from Neanderthals and modern humans. Sequencing revealed that the individual belonged to a distinct species of human, named by the discoverers as’Denisovans’ after the cave in which the first fossil remains were found, and that this new species of humans split from Neanderthal populations about 381 – 473 kyr ago.[3]
The Denisovans were apparently not a blip on the screen of human evolution. Recent sequencing of the mitochondrial genome of a roughly 400,000-year-old hominin fossil recovered from the Sima de los Huesos cave in Spain revealed that it belonged to a hominin from a population characterized as a sister group to Denisovans [4]; and genetic evidence of admixture with Denisovans between anatomically modern humans (AMH) has shown up in Melanesian groups, especially those of Papua New Guinea, and to a lesser extent among mainland Asians and Native Americans.
Who were these Denisovans and what did they look like?
We know what Neanderthals looked like, based on the fossil record. But the fossil record for Denisovans has been very paltry by comparison. Here again, genetic sequencing and analysis has made a difference. Another landmark genome sequencing study, this time in 2012, was conducted on the Denisovan finger bone by a team led by Svante Pääbo of the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany, revealing a young girl with dark skin, brown hair, and brown eyes—the sequence indicated genetic variants that in present-day humans are associated with these physical traits. Moreover, from this study the researchers were able to determine that the genetic variation of the Denisovans was lower than in present-day humans, and suggested that an initially small Denisovan population grew quickly and spread over a wide geographic range. Coupled with similar genetic research on Neanderthals, the results even spelled possible implications for the early dispersal of hominins from their African homeland. “If future research of the Neanderthal genome shows that their population size changed over time in similar ways,” said Pääbo, “it may well be that a single population expanding out of Africa gave rise to both the Denisovans and the Neanderthals.” The researchers generated a list of about 100,000 changes in the human genome that occurred after the split from the Denisovans. Some of the changes are associated with brain function and nervous system development. Others may be related to the skin, eye and dental morphology.
“This research will help determining how it was that modern human populations came to expand dramatically in size as well as cultural complexity while archaic humans [such as the Denisovans and Neanderthals] eventually dwindled in numbers and became physically extinct”, said Pääbo.
The Denisovan discovery was another reminder to evolutionists that the landscape of human evolution was far more complicated than what had previously been imagined.
________________________________________
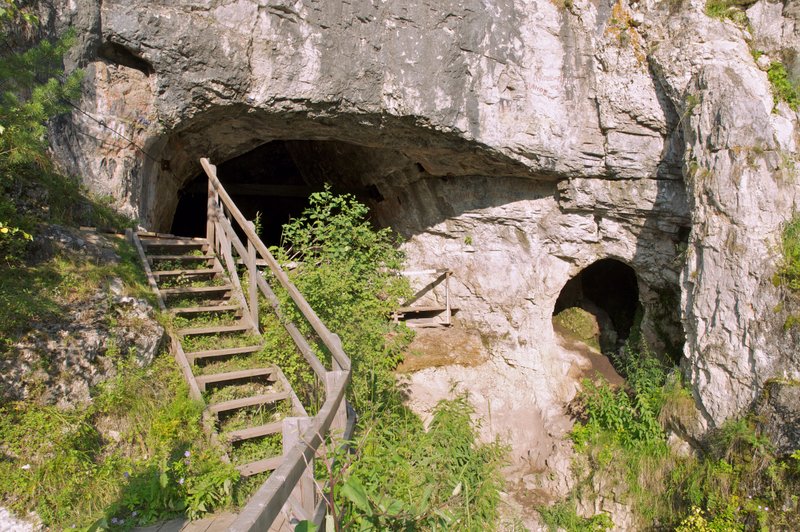 This is the Denisova Cave entrance, located in the Altai Mountains of southern Siberia, Russia. The cave was inhabited at various times by three different groups of early humans: Neanderthals, Denisovans and modern humans. Credit: Bence Viola
This is the Denisova Cave entrance, located in the Altai Mountains of southern Siberia, Russia. The cave was inhabited at various times by three different groups of early humans: Neanderthals, Denisovans and modern humans. Credit: Bence Viola
________________________________________
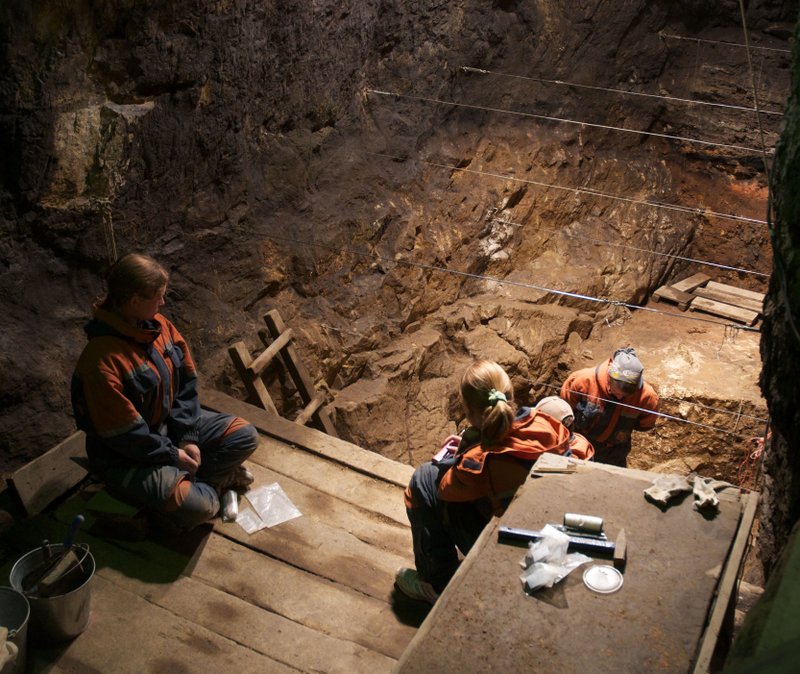 Excavation shown in progress inside the Denisova cave. This image relates to a paper that appeared in the Aug. 31, 2012, issue of Science, published by AAAS. The paper, by M. Meyer at Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany, and colleagues was titled, “A High-Coverage Genome Sequence from an Archaic Denisovan Individual.” Image courtesy of Max Planck Institute for Evolutionary Anthropology
Excavation shown in progress inside the Denisova cave. This image relates to a paper that appeared in the Aug. 31, 2012, issue of Science, published by AAAS. The paper, by M. Meyer at Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany, and colleagues was titled, “A High-Coverage Genome Sequence from an Archaic Denisovan Individual.” Image courtesy of Max Planck Institute for Evolutionary Anthropology
________________________________________
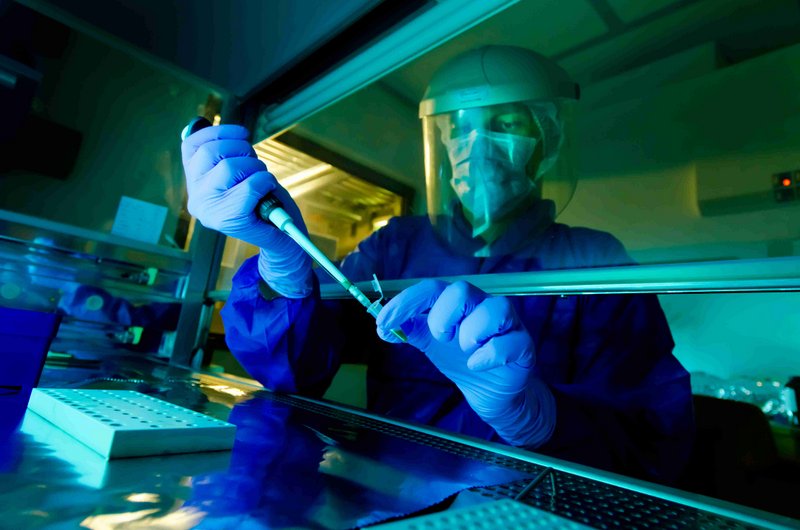 In the genetics lab: Matthias Meyer at work in the clean lab. Meyer was co-author of the 2012 Denisova genome study. Credit:MPI for Evolutionary Anthropology
In the genetics lab: Matthias Meyer at work in the clean lab. Meyer was co-author of the 2012 Denisova genome study. Credit:MPI for Evolutionary Anthropology
_____________________________________
Mixing it Up
The possibility of admixture, or interbreeding, between AMHs and archaic humans such as Neanderthals has been the subject of anthropological discussion and debate for decades. Finally, the recent characterization of genomes from archaic human remains has provided convincing evidence that such events occurred. The more abundant findings have related to the relationship between Neanderthals and AMHs.
For example, one recent study [5] by a University of California, Berkeley, research team achieved a complete sequence of the Neanderthal genome using DNA extracted from a woman’s toe bone (also excavated in Denisova Cave in the Altai Mountains) dating back to 50,000 years, revealing a history of interbreeding among at least four different types of early humans living in Europe and Asia at that time. In this study, they generated a high-quality sequence of the Neanderthal genome and compared it with the genomes of modern humans and Denisovans. Their comparison showed that Neanderthals and Denisovans were closely related, and that their common ancestor split from the ancestors of modern humans about 400,000 years ago. The study also showed that Neanderthals and Denisovans split about 300,000 years ago. The study indicated that, though Denisovans and Neanderthals became extinct, they left behind traces of their genes because they occasionally interbred with AMHs. They estimated that between 1.5 and 2.1 percent of the genomes of modern non-Africans can be traced to Neantherthals. Denisovan genetic traces were also found in some Oceanic and Asian modern human populations. The genomes of Australian aborigines, New Guineans and some Pacific Islanders were determined to be about 6 percent Denisovan genes, according to earlier studies. The new analysis found that the genomes of Han Chinese and other mainland Asian populations, as well as of Native Americans, contain about 0.2 percent Denisovan genes. More interesting still was the finding that Denisovans interbred with a mysterious fourth group of early humans also living in Eurasia at the time. That group split away from the others more than a million years ago, and scientists suggest this group may have been Homo erectus, the archaeological and fossil evidence of which has indicated their presence in Europe and Asia a million or more years ago.
“The paper really shows that the history of humans and hominins during this period was very complicated,” said population geneticist Montgomery Slatkin, a study co-author and a UC Berkeley professor of integrative biology. “There was a lot of interbreeding that we know about and probably other interbreeding we haven’t yet discovered.” Although interbreeding, based on the study results, is thought to have been relatively infrequent between Neanderthals and AMHs.
In another analysis, they discovered that the Neanderthal woman whose toe bone provided the DNA was highly inbred. The woman’s genome indicated that she was the daughter of a mother and father who either were half-siblings who shared the same mother, an uncle and niece or aunt and nephew, a grandparent and grandchild, or double first-cousins (the offspring of two siblings who married siblings). Additional analyses suggested that the population sizes of Neanderthals and Denisovans were small and that inbreeding may have been more common in Neanderthal groups than in modern populations. Moreover, they identified at least 87 genes in modern humans that differ significantly from corresponding genes in Neanderthals and Denisovans. These genes may hold key clues to the behavioral differences between modern humans and the extinct, archaic human species. According to Svante Pääbo of the Max Planck Institute, they could constitute “a catalog of genetic features that sets all modern humans apart from all other organisms, living or extinct.”
“I believe,” he added, “that in it hide some of the things that made the enormous expansion of human populations and human culture and technology in the last 100,000 years possible.”
In another new study, conducted by researchers under the leadership of Benjamin Vernot and Joshua Akey of the University of Washington, Seattle, researchers came up with a proven methodology to determine the percentages and precise genome segments that have been inherited, and which parts of the total Neanderthal genome bestowed an adaptive advantage (and were thus retained) in modern human descendants. [6] Their model involved a two-staged computational strategy framework, without additional sampling of fossil remains, applied to whole-genome sequences of 379 Europeans and 286 East Asians, courtesy of data from the 1000 Genomes Project. The results of their research suggested that, while the total amount of Neanderthal sequence in any individual modern human is relatively low, about 2 – 4 percent, the cumulative amount of the Neanderthal genome identified across all humans in the aggregate represents segments that constituted about 20 percent of a total Neanderthal genome sequence. In some parts of the modern genome, they observed large regions without Neanderthal DNA, suggesting that certain portions of the archaic genetic sequence were deleterious to survival. On the other hand, they also observed sections of the modern sequence that showed more Neanderthal DNA than expected. Vernot and Akey concluded that these sequences remained because their functions provided an adaptive advantage, perhaps related to skin phenotype.
Their study could have implications for future research.
“Our results provide a new avenue for paleogenomics studies,” wrote Vernot and Akey in their report, “allowing substantial amounts of population-level DNA sequence information to be obtained from extinct groups even in the absence of fossilized remains……. potentially allowing the discovery and characterization of previously unknown hominins that interbred with modern humans.”[6]
In a related study published in the Jan. 29 issue of the journal Nature, another team of scientists led by Harvard Medical School geneticist David Reich, including Svante Pääbo of the Max Planck Institute, obtained results suggesting that modern human populations inherited other genetic traits from Neanderthals that are connected to both positive adaptive functions and characteristics and those that could be described as negative. They found traces of Neanderthal DNA, for example, that affect the keratin filaments in the skin—proteins that make hair, nails and skin tougher for surviving colder climates; genes that affect the immune system; and genes that affect such conditions as Crohn’s disease, type 2 diabetes, smoking behavior, billiary cirrhosis, and lupus. The researchers did this by analyzing the genetic variants found in 846 non-African people and 176 people from sub-Saharan Africa, and then compared the results to that of a 50,000-year-old Neanderthal with a relatively intact genome sequence.
Finally, in a yet more recent study published in the journal Nature, Svante Pääbo of the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany and Howard Hughes Medical Institute (HHMI) investigator David Reich at Harvard Medical School, along with researchers at the Key Laboratory of Vertebrate Evolution and Human Origins in Beijing, China, were able to tease out and analyze a small percentage of the DNA remnants found in a 37,000 to 42,000-year-old modern human jaw bone originally found in 2002 by cavers and archaeologists in the Oase Cave in south-western Romania. The fossil itself has been considered evidence of the oldest known modern human in Europe.
The results of their study were astonishing. By comparing the fossil’s genome to genetic data from other groups and then conducting a series of statistical analyses, they determined that the human to which the fossil jaw bone belonged had a Neanderthal ancestor removed by only 4 to 6 generations back.
“I could hardly believe it when we first saw the results,” said Pääbo. “It is such a lucky and unexpected thing to get DNA from a person who was so closely related to a Neanderthal.”
“The sample is more closely related to Neanderthals than any other modern human we’ve ever looked at before,” Reich added. “We estimate that six to nine percent of its genome is from Neanderthals. This is an unprecedented amount. Europeans and East Asians today have more like two percent.”
Thus, not only was the Oase specimen the oldest modern human in Europe known so far, but now research confirmed that the individual had a relatively closely connected Neanderthal ancestor, further evidence that at least some Neanderthals and modern humans, for a time, comingled at a level something beyond a hand shake or exchange of goods.
The evidence for admixture and the affects of it will likely continue to emerge as scientists further develop techniques and technologies in genetic analysis and as new finds are uncovered through archaeological and other field research.
____________________________________________
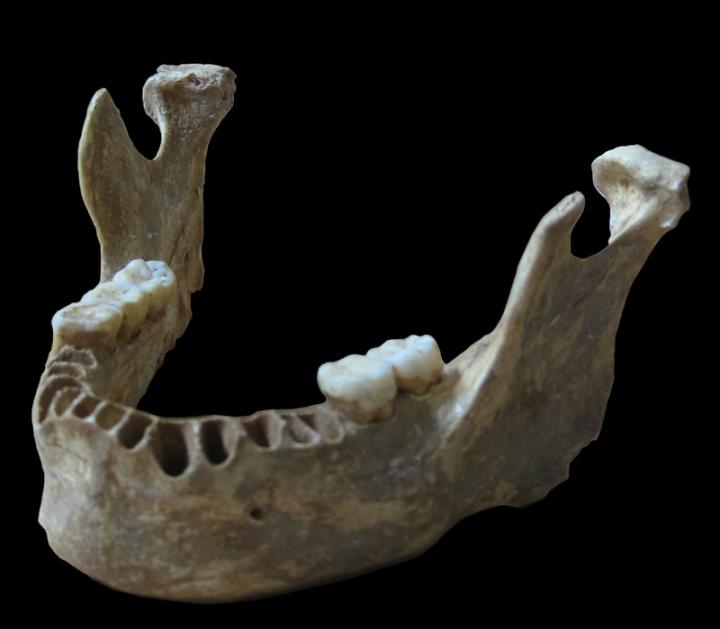 DNA taken from a 40,000-year-old modern human jawbone reveals that this man had a Neandertal ancestor as recently as four to six generations back. Courtesy MPI f. Evolutionary Anthropology/Pääbo
DNA taken from a 40,000-year-old modern human jawbone reveals that this man had a Neandertal ancestor as recently as four to six generations back. Courtesy MPI f. Evolutionary Anthropology/Pääbo
__________________________________________________________
Out of Africa
According to the preponderance of aDNA study results, anatomically modern humans originated in Africa. The big debate has revolved around when, how, and the tempo of dispersal out of Africa into Eurasia and further southeast into mainland East Asia and Melanesia/Australia. Generally, the suggested exodus date has been estimated to be between 50 and 62 kyr ago, branching from a small population of about 1,000 individuals. From this point, two contending models have topped the list of proposed dispersal paradigms.
The first, called the ‘single dispersal’ hypothesis, suggests an initial dispersal about 50 kyr ago out of Africa into Eurasia, supported by genetic studies that show descendants of all non-African lineages from a single ancestor who lived around 55-75 kyr ago. The initial single dispersal was followed by a series of ‘founder’ and separate migrations into Asia (55-40 kyr ago) and Europe (40-25 kyr ago), with a further expansion from Asia into Australia as early as 50-40 kyr ago as the ancestral group to Aboriginal Australians.
The second model, called the ‘multiple dispersal’ hypothesis, suggests separate successive migrations out of Africa, including a very early dispersal that accounts for an ancestral Aboriginal Australian population at about 60-50 kyr ago. Interesting for this hypothesis was the sequencing of the complete genome of an ancient Aboriginal Australian taken from a hair tuft. It indicated that the ancestors of Aboriginal Australians spread into East Asia around 62-75 kyr ago, interbreeding with Denisovans along the way and arriving in Australia about 50 kyr ago. The model further suggests that the ancestors of present-day mainland Asians followed in a second, independent wave at around 38-25 kyr ago.
Most recently, a genomic study conducted by Dr. Luca Pagani of the Wellcome Trust Sanger Institute and the University of Cambridge in the UK and colleagues suggested that modern humans made their first successful major migration out of Africa around 55,000 – 60,000 years ago through Egypt. The results of that study has informed the ongoing scholarly debate related to whether the first major modern human migration made its way initially out of Africa by taking a northern route through Egypt, or by traveling from a more southerly orientation northward through Ethiopia and up through the Arabian Peninsula.
“In our research,” explained Pagani, “we generated the first comprehensive set of unbiased genomic data from Northeast Africans and observed, after controlling for recent migrations, a higher genetic similarity between Egyptians and Eurasians than between Ethiopians and Eurasians.”
Pagani’s research team accomplished this by analyzing genetic information from six modern Northeast African populations, consisting of 100 Egyptians and five Ethiopian populations, each represented by 25 people. The team also used high-quality genomes to estimate the time that the populations split from one another: the genomes of people outside Africa showed a split from the Egyptian genomes more recently than the split from Ethiopians (55,000 as opposed to 65, 000 years ago), supporting the idea that Egypt was the last stop on the route out of Africa.
“This important study still leaves questions to answer,” says Dr Chris Tyler-Smith, a senior author from the Wellcome Trust Sanger Institute. “For example, did other migrations also leave Africa around this time, but leave no trace in present-day genomes? To answer this, we need ancient genomes from populations along the possible routes. Similarly, by adding present-day genomes from Oceania, we can discover whether or not there was a separate, perhaps Southern, migration to these regions.”
Needless to say, a complete and accurate picture of AMH dispersal may be a long way off, although future genetic studies, coupled with refinements and innovations in genetic sequencing and analysis of ancient human remains (if recent history of the science is any indicator) show exciting promise.
__________________________________________
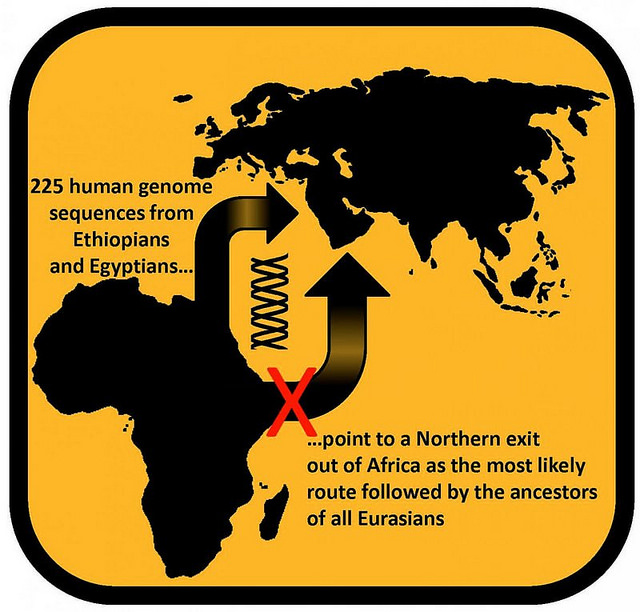 Human genome sequences from Ethiopians and Egyptians point to a Northern exit out of Africa as the most likely route by the ancestors of all Eurasians. Image courtesy Luca Pagani
Human genome sequences from Ethiopians and Egyptians point to a Northern exit out of Africa as the most likely route by the ancestors of all Eurasians. Image courtesy Luca Pagani
________________________________________________________
The First Americans
Hotly debated among researchers have been the questions of when, how and where the first peopling of the Americas occurred. Recent archaeological research has cast new light and raised new questions about who the ‘first Americans’ were and when and how they entered the American continents, not the least of which have been the recent archaeological findings at sites such as the Paisley Caves in Oregon, the Manis site in Wisconsin, the Anzick site in Montana, and the Meadowcroft rockshelter in Pennsylvania, to name only a few. These sites, among others, have arguably pushed back the clock on the timing of the first peopling, challenging the long-held tradition that the Clovis culture represented the first population to enter the Americas. The broadly accepted view about when and how people first entered the Americas has revolved in part around the changes in the glacial periods associated with the last glacial period of the Ice Age, a time when the Cordilleran and Laurentide ice sheets covered much of northern North America. However, during the warmer interglacial periods, they retreated to create ice-free corridors along the Pacific coast and areas east of the Rocky Mountains of Canada. Scientists have long suggested that it was through these corridors that humans were likely able to cross Beringia into the Americas. Beringia was a land bridge as much as 1,000 miles wide that joined present-day Alaska and eastern Siberia at various times 110,000 to 10,000 years ago. Exactly when and how this crossing may have occurred has been open to question for decades, but traditionally the most widely accepted proposal advanced the suggestion that it may have occurred around 15,000 years ago, giving rise to the Clovis culture—until the discovery of pre-Clovis type artifacts at sites such as those mentioned above.
DNA evidence for earlier settlement of the Americas has continued to mount. DNA was found and analyzed in human coprolites excavated at Paisley Caves in south-central Oregon, dating to approximately 12.3 kyr ago.[7] DNA analyses of the tip of a bone projectile point implanted in the rib bone of a disarticulated mastodon uncovered at the Manis site in Wisconsin and dated to about 11.9 kyr ago, brought additional evidence that humans already hunted such large mammals very early on, further supporting a pre-Clovis settlement of the Americas.[8] The genome of a 24 kyr old individual from the Mal’ta site in south-central Siberia (Russian Federation) was sequenced, revealing a new window on the origins of Native Americans outside of the American continents.[9] A Mal’ta Y chromosome haplotype corresponded to a basal lineage of haplogroup R, which is widespread in modern-day western Eurasians and is thought to be a sister lineage to ‘haplogroup Q’, considered the most common haplogroup in Native Americans. Scientists suggest that this gene flow occurred prior to diversification of Native American populations in the Americas, possibly representing 14—38% of Native American ancestry. The western Eurasian ancestry finds suggest that Native Americans may have derived the European component not only from post-Columbian interbreeding, but from an earlier mixed ancestry that included a European contribution.
________________________________________
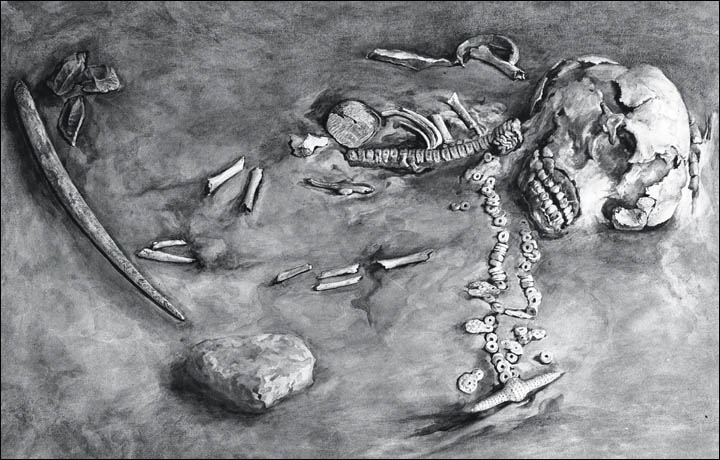 The remains of the 24,000-year-old Mal’ta boy. Credit: State Hermitage Museum in Russia
The remains of the 24,000-year-old Mal’ta boy. Credit: State Hermitage Museum in Russia
_________________________________________
But add to this the sequencing of a complete genome of a 12.6 kyr old child who was buried among ochre-covered Clovis artifacts at the Anzick site Montana.[10] This genome showed a much greater genetic affinity with Native Americans, particularly Central and South Americans, than with any extant Eurasian population or North American Native populations. It suggested that the North American and South American lineages diverged early, the Anzick individual being related to the South American lineage, possibly ancestral to all South American populations, or possibly ancestral to all Native Americans. This individual used Clovis tools. Many scientists theorize that since both the Anzick and Mal’ta individuals shared affinities to both western and eastern Eurasians, gene flow from the Mal’ta population of Siberia into Native American population groups likely happened before about 12.6 kyr ago.
And finally, initially discovered in 2007 by a team of underwater archaeologists, a nearly complete skeleton was found within a submerged chamber (which site investigators called “Hoyo Negro”, or “black hole”) in the Sac Actun cave system on Mexico’s Eastern Yucatán Peninsula. Named “Naia” by the dive team, she was, according to forensic analysis, a slightly-built teenage girl at death, measuring only 4 feet 10 inches in height. Anthropologists who examined the skeleton determined that she was between 15 and 16 years old, and had likely fallen to her death into the chamber (a dry pit during the time of the fall) before the cave had subsequently filled with water due to climate change and rising sea levels.
The find was unprecedented for two reasons. The first reason had to do with the age. Scientists dated the skeleton to between 12,000 and 13,000 years ago based on radiocarbon dating of tooth enamel and Uranium/Thorium dating analyses of mineral deposits on her bones. The second reason was related to the pristine preservation of the bones, which facilitated the study team’s analysis of the girl’s mitochondrial DNA (mtDNA), a useful tool for examining the relatedness of populations. Their analysis revealed a haplotype common to modern Native Americans, subhaplogroup D1. This genetic signature occurs only in the Americas. The report authors suggested that this genetic signature likely developed in Beringia after populations there split from other Asians, indicating that individuals of this population traveled at least far and wide enough through the Americas to have reached Mexico by 12,000 – 13,000 BP.
But there was one fly in the ointment. The cranio-facial features of this skeleton differed noticeably form those of most Native Americans today. To this the researchers suggested that the differences in craniofacial form were probably best explained as evolutionary changes that happened after the divergence of Beringians from their Siberian ancestors. Thus the Americas, they theorize, were not colonized by separate migration events from different parts of Eurasia. Rather, the earliest Americans represent an early population expansion out of Beringia. This aligns with the hypothesis that both Paleoamericans and Native Americans derive from a single source population, hunter-gatherers who moved onto the Bering Land Bridge from northeast Asia (Beringia) between 26,000 and 18,000 years ago, spreading southward into North America sometime after 17,000 years ago. [11]
______________________________________
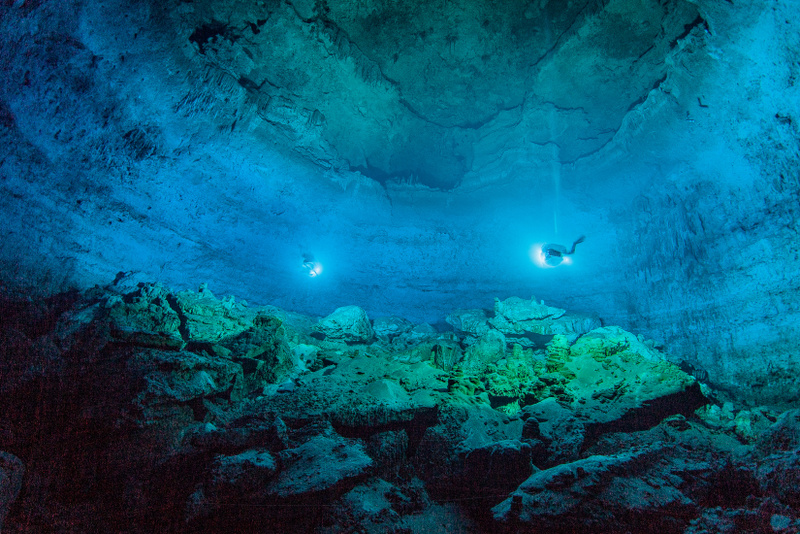 A broad view of Hoyo Negro, shot from the floor near the south edge, showing the immensity of the chamber and the complexity of the boulder-strewn bottom. One access tunnel can be seen near the ceiling at top left. This photo was taken by the “painting with light” method on a 30 second exposure. Text and photo credit: Roberto Chavez Arce
A broad view of Hoyo Negro, shot from the floor near the south edge, showing the immensity of the chamber and the complexity of the boulder-strewn bottom. One access tunnel can be seen near the ceiling at top left. This photo was taken by the “painting with light” method on a 30 second exposure. Text and photo credit: Roberto Chavez Arce
_______________________________________
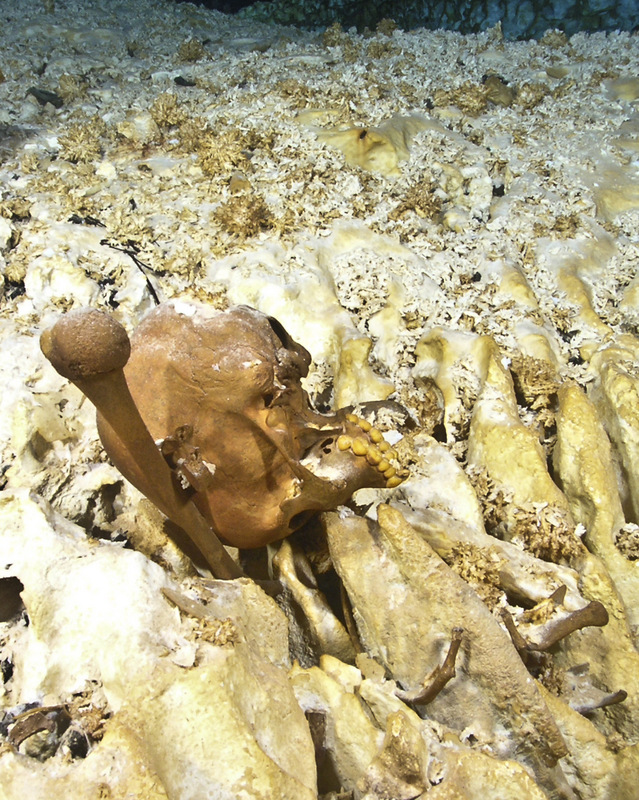 The skull of Naia as it was discovered in 2007, resting against the left humerus (upper arm bone).
The skull of Naia as it was discovered in 2007, resting against the left humerus (upper arm bone).
Photo and text credit: Daniel Riordan Araujo
_______________________________________
Research, genetic and otherwise, continues on the question of the first Americans. More surprises may be in store.
Our Big Brains
Without question, the human brain has been perhaps the single-most distinctive morphological and functional characteristic that has separated our species from the rest of the primates. Here, too, genetic research is shedding new light on the role and significance this organ has played in our evolutionary past, and what makes it unique among the planet’s biological fabric.
In a recent study, for example, Duke University researchers mined databases of genomic data from humans and chimpanzees, finding ‘enhancers’ that were present in the brain tissue.[12] They studied the enhancers that significantly differed between the two primate species, identifying genes key to brain development. They called them ‘human-accelerated regulatory enhancers,’ or HARE, connected to the proliferation of new neurons. One particular enhancer stood out, which they called ‘HARE5’. They compared the HARE5 to its equivalent in the chimpanzee by expressing them into mice embryos. What they found was that, though the human HARE5 and the chimpanzee HARE5 sequences differed by only 16 letters in their genetic code, the human enhancer was active earlier in brain development and more active generally than the chimpanzee enhancer. The human HARE5 mice embryos developed brains 12% larger in area compared with chimpanzee HARE5 mice to gestation. The growth centered on the neocortex, the region of the brain involved in higher-level function such as language and reasoning.
“What we found is a piece of the genetic basis for why we have a bigger brain,” said study co-author Gregory Wray, professor of biology and director of the Duke Center for Genomic and Computational Biology. “It really shows in sharp relief just how complicated those changes must have been.”
________________________________________
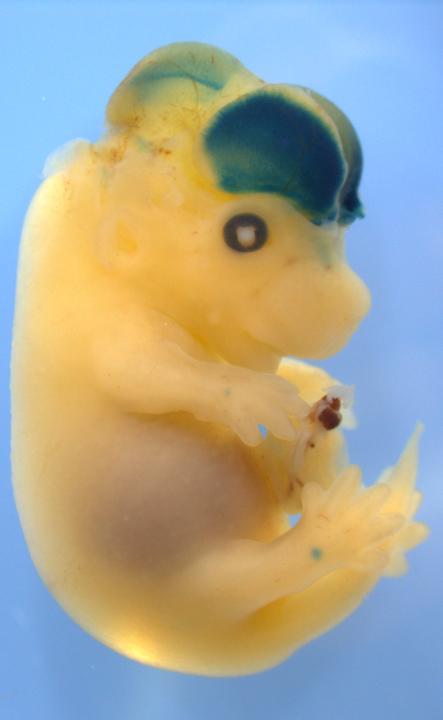 The human version of a DNA sequence called HARE5 turns on a gene important for brain development (gene activity is stained blue), and causes a mouse embryo to grow a 12 percent larger brain by the end of pregnancy than an embryo injected with the chimpanzee version of HARE5. Image Credit: Silver lab, Duke University
The human version of a DNA sequence called HARE5 turns on a gene important for brain development (gene activity is stained blue), and causes a mouse embryo to grow a 12 percent larger brain by the end of pregnancy than an embryo injected with the chimpanzee version of HARE5. Image Credit: Silver lab, Duke University
______________________________________
In another similar recent study, a research team headed by Marta Florio of the Max Planck Institute of Molecular Cell Biology and Genetics identified a gene that, like the HARE5 gene regulatory enhancer in the Duke University study, drives the proliferation of neural progenitor cells in the neocortex.[13] The gene, they maintain, is common among modern humans, Neanderthals, and Denisovans, but not other primates. Called ‘ARHGAP11B’, it is a hominin-specific gene. The researchers suggested that it arose in the human lineage soon after humans diverged from chimpanzees. Like the Duke University study, when Florio and her colleagues expressed the gene into a developing mouse brain, they found that a key section of the neocortex grew significantly larger than what would be expected in a normal mouse.
______________________________________
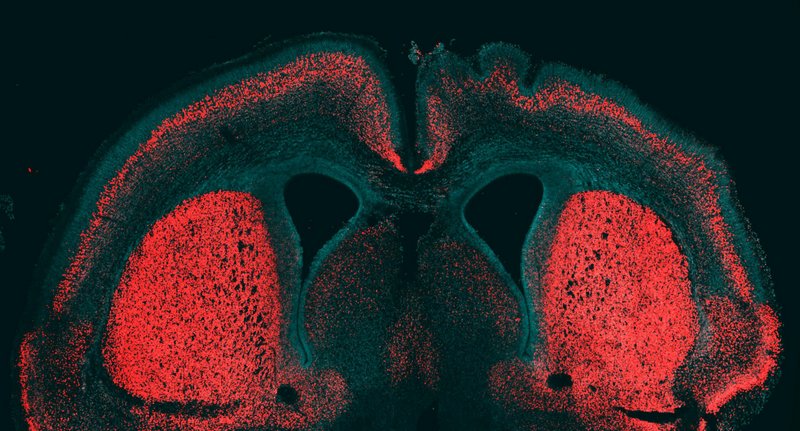 Embryonic mouse cerebral cortex stained for cell nuclei and a marker of deep-layer neurons (red). The human gene ARHGAP11B was expressed into the right hemisphere. Credit: Marta Florio and Wieland B. Huttner, Max Planck Institute of Molecular Cell Biology and Genetics
Embryonic mouse cerebral cortex stained for cell nuclei and a marker of deep-layer neurons (red). The human gene ARHGAP11B was expressed into the right hemisphere. Credit: Marta Florio and Wieland B. Huttner, Max Planck Institute of Molecular Cell Biology and Genetics
______________________________________
What were the factors that influenced this genetic change in human/primate evolutionary history? That is the ‘million dollar question’, so to speak. It has, and will continue, to require a global, multidisciplinary effort by scientists to come up with the answers.
From Mesolithic to Neolithic in Europe
A key debate in more recent human evolutionary studies has revolved around understanding the genetic history of Europeans, and central to the discussion has been the impact of the Neolithic transition (the switch from hunting-gathering to farming) on the European gene pool. According to supporters of what has been called the ‘demic diffusion’ model, the Neolithic transition in Europe was defined at least in part by migrations of early farmers from the Near East, suggesting a genetic ‘discontinuity’ between pre-Neolithic, or Mesolithic, populations and Neolithic and post-Neolithic populations. Under this model, a significant Near Eastern Neolithic ancestry should show up in the current gene pool. Conversely, the ‘cultural diffusion’ model suggests the diffusion of ideas and technology, and not people, which would result in a genetic continuity between early farmers and hunter-gatherers and thus a much less significant Near Eastern marker in the gene pool.
These models have been genetically tested on a variety of European remains. Not altogether surprising to many theorists, the genetic data from post-Mesolithic prehistoric individuals has generally shown that the Neolithic was the transition primarily responsible for a genetic discontinuity between hunter-gatherers and present-day populations, and that there is a genetic affinity between Early Neolithic individuals and present-day Near Eastern populations. In other words, much of the data has supported the demic diffusion model.
It is more complicated than this, however. The data also suggested some genetic discontinuity between Early Neolithic people and today’s Europeans, pointing to possible later population movements. For example, genomic data from the now famous Tyrolean ‘Iceman’, dated to about 5,000 year ago, indicated a genetic affinity with the Funnel Beaker Culture, a hunter-gatherer population of Scandinavia. This suggested that these Scandinavian groups were geographically widespread at that time. [14]
Genetic studies conducted on samples outside of ancient human remains have also shed light and raised new questions about this period of time in Europe, further complicating the emerging picture. One recent study serves to illustrate this point well:
In an analysis of sedimentary ancient DNA from an underwater site off the southern coast of Britain, a U.K research team found that Mesolithic hunter-gatherers who occupied a site now underwater traded in a Near Eastern strain of wheat 2,000 years before the currently generally-accepted advent of farming in Britain. [15]
“The first evidence of cereal cultivation on what is now mainland Britain dates back only to about 6,000 years ago, suggesting a substantial temporal gap between the two sides of the English Channel,” wrote University of Oxford’s Greger Larson in a perspective article published about the finding in Science magazine. Larson is Director of the Palaeogenomics & Bio-Archaeology Research Network Research Laboratory for Archaeology and the History of Art. It is generally thought that farming began in Europe, south and west of the English Channel, much earlier than in the British Isles. He went on to summarize the process and merits of the recently completed study by the team of U.K. scientists led by Oliver Smith of the University of Warwick, which suggested the presence of not just wheat, but a Near Eastern strain of wheat, within 8,000-year-old submerged paleosol sediments at Bouldnor Cliff off the Isle of Wight in the English Channel.
Smith and colleagues teased DNA from core samples taken from a Mesolithic paleosol layer just beneath a peat layer sealed beneath silty-clay submerged alluvial sediments. Millennia ago, this paleosol layer was above ground, an ancient landscape that was gradually submerged as sea levels rose during the warming period beginning in the early Holocene epoch. The sea level change inundated the land bridge between what is today Britain and the rest of Europe, creating the English Channel. The researchers uncovered evidence of human occupation typical of Mesolithic hunter-gatherers at the site, which included worked wood, burnt flint and hazelnut shells.
“The site has been dated to 8030 to 7980 calendar (cal) yr B.P.,” wrote Smith, et al. in their report, “placing it in the late Mesolithic of the British Isles, a period that is represented by few assemblages and is still little understood………The sedaDNA [sedimentary ancient DNA] profile revealed a wooded landscape that included oak, poplar, apple, and beech family members, with grasses and a few herbs present.”
But the most startling finding came from the DNA analysis. Through meticulous analysis accounting for and dismissing possible contamination as well as potential intrusion from other upper layers, they discovered clear DNA traces of Near Eastern strains of wheat genuinely dated to and associated within the context of the Mesolithic assemblage, which also included evidence of a Mesolithic human diet.
“The occurrence of wheat 8,000 years ago on the British continental shelf appears early, given its later establishment on the UK mainland,” wrote Smith, et al., “Neolithic assemblages first appear in northwest Europe in the 8th millennium B.P., from 7500 B.P. in the central Rhineland, 7300 B.P. in the Rhine/Maas delta and adjacent areas, and 7400 B.P. in western France.”
The DNA results thus suggested the presence of wheat at Bouldnor Cliff about 400 years before the earliest known occurrences of farming in northwestern Europe, and 2,000 years before agriculture is known to have taken hold in what is today Britain.
The researchers found no evidence that the wheat had actually been cultivated at or near the site. Instead, they suggested the wheat was likely traded into the area by a network established between hunter-gatherers at Bouldnor Cliff and Neolithic farmers further south and west in Europe.
“We suspect that this wheat represents foodstuffs imported from the continent rather than the cultivation of this cereal crop at this locale. The presence of wheat, along with pioneering technological artifacts at the site, provides evidence for a social network between well-developed Mesolithic peoples of northwest Europe and the advancing Neolithic front,” concluded Smith, et al., suggesting the agricultural products moved ahead of their actual cultivation in Britain.
The point of the foregoing example is that, although the predominant thinking among scholars has placed the advent of the farming Neolithic in what is today Britain by 6000 years BPE, the timing and mode of Neolithization is still debated. As such, this study has important implications in the ongoing research on the evolution of agriculture in Europe.
“The unexpectedly early appearance of wheat in Britain should force a rethinking of both the strength of the relationships between early farmers and hunter-gatherers, and the origins of settled agricultural communities in Europe,” concluded Larson in the Science perspective article.
________________________________________
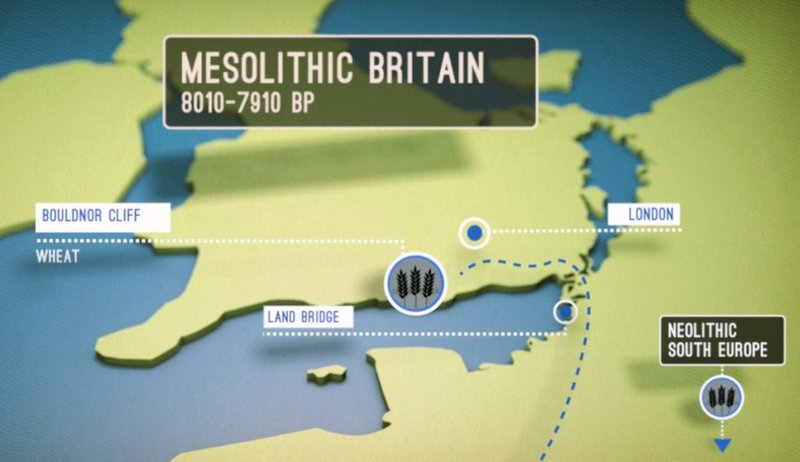 Above: From the Youtube video (screenshot) describing the research findings in the paper titled “Sedimentary DNA from a submerged site reveals wheat in the British Isles 8,000 years ago”. Credit: University of Warwick
Above: From the Youtube video (screenshot) describing the research findings in the paper titled “Sedimentary DNA from a submerged site reveals wheat in the British Isles 8,000 years ago”. Credit: University of Warwick
_________________________________________
The First Great Population Explosion?
The prevailing theory regarding human population expansion revolves around the generally accepted notion that human populations rapidly increased with the advent of agriculture. The archaeological record bears this out. But one interesting genetic study has provided evidence of another, earlier “population explosion” before modern humans settled into communities centered around agriculture and animal husbandry:
Carla Aimé and her colleagues at Laboratoire Eco-Anthropologie et Ethnobiologie, University of Paris, conducted a study using 20 different genomic regions and mitochondrial DNA of individuals from 66 African and Eurasian populations, and compared the genetic results with archaeological findings. [16] They concluded that the first big expansion of human populations may be much older than the one associated with the emergence of farming and herding, and that it could date as far back as Paleolithic times, or 60,000-80,000 years ago. The humans who lived during this time period were hunter-gatherers. The authors hypothesized that the early population expansion could be associated with the emergence of new, more sophisticated hunting technologies, as evidenced in some archaeological findings. Moreover, they state, environmental changes could possibly have played a role.
The researchers also showed that populations who adopted the farming lifestyle during the Neolithic Period (10,200 – 3,000 B.C.) had experienced the most robust Paleolithic expansions prior to the transition to agriculture.
“Human populations could have started to increase in Paleolithic times, and strong Paleolithic expansions in some populations may have ultimately favored their shift toward agriculture during the Neolithic,” said Aimé.
The details of the study have been published in the scientific journal, Molecular Biology and Evolution, by Oxford University Press.
The Promise of the Future
Genetic research has already provided a massive amount of new data that has in many ways revolutionized what we know about our prehistoric past. And though this writing has only touched the surface of what has been accomplished in the last two decades, it has hopefully afforded an appetizing glimpse for readers of what has been happening. To date, we have genomic data on at least 24 individuals, of which 8 are archaic humans, that have informed our understanding of human evolution. That number will no doubt increase. At this writing, new advances are under development that will push back the human prehistoric and evolutionary timeline available for examination, including a sharpening of our focus and discovery of new events related to what was going on in our ancestral past. With the ongoing development of such new things as ‘high-throughput sequencing’ technologies, as well as technologies capable of targeting ultra-short and ultra-damaged ancient DNA fragments and, beyond genetics, the development of ancient protein sequencing techniques, scientists will make it possible for us to see far deeper into our prehistoric past with a clarity likely unachievable through the archaeological and paleontological record alone.
______________________________________________________
[1]* Templeton, Alan R., Revolutionizing the ‘Out of Africa’ Story, Genetic Engineering and Biotechnology News, April 1, 2013, Vol 33, No. 7 (online)
** Genomic and cranial phenotype data support multiple modern human dispersals from Africa and a southern route into Asia,” by Hugo Reyes-Centeno et al. http://www.pnas.org/cgi/doi/10.1073/pnas.1323666111
[2] Green, R.E., A draft sequence of the Neandertal genome, Science 328, 71 – 722.
[3] Krause, J., et al., The complete mitochondrial DNA genome of an unknown hominin from southern Siberia, Nature 464, 894 -897.
[4] Meyer, M., et al., A mitochondrial genome sequence of a hominin fro Sima de los Huesos. Nature 505, 403 – 406.
[5] Prufer, Kay, et al., The complete genome sequence of a Neanderthal from the Altai Mountains, Nature, Vol. 505, Issue 7481, 02 January 2014.
[6 ] “Resurrecting Surviving Neanderthal Lineages from Modern Human Genomes,” by B. Vernot; J.M. Akey at University of Washington in Seattle, WA, Science, 29 January, 2014.
[7] Gilbert, M.T., et al., DNA from pre-Clovis human coprolites in Oregon, North America. Science 320, 786 – 789.
[8] Waters, M.R., Pre-Clovis mastodon hunting 13,800 years ago at the Manis site, Washington. Science 334, 351 – 353.
[9] Raghaven, M., et al., Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505 87 – 91.
[10] Rasmussen, M., et al., The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 506, 225 – 229.
[13] https://popular-archaeology.com/issue/winter-01012015/article/a-brain0building-gene-unique-to-humans
[14] Skoglund, P., et al., Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466 – 469. – and – Skoglund, P., et al., Genomic diversity and admixture differs for Stone-Age Scandinavian foragers and famers. Science 344, 747 – 750.
Ermini, L., et al., Major transitions in human evolution revisited: A tribute to ancient DNA, Journal of Human Evolution (2014), http://dx.doi.org/10.1016/j.jhevol.2014.06.015
______________________________________________________





