HARVARD MEDICAL SCHOOL—The introduction of agriculture into Europe about 8,500 years ago changed the way people lived right down to their DNA.
Until recently, scientists could try to understand the way humans adapted genetically to changes that occurred thousands of years ago only by looking at DNA variation in today’s populations. But our modern genomes contain mere echoes of the past that can’t be connected to specific events.
Now, an international team reports in Nature that researchers can see how natural selection happened by analyzing ancient human DNA.
“It allows us to put a time and date on selection and to directly associate selection with specific environmental changes, in this case the development of agriculture and the expansion of the first farmers into Europe,” said Iain Mathieson, a research fellow in genetics at Harvard Medical School and first author of the study.
By taking advantage of better DNA extraction techniques and amassing what is to date the largest collection of genome-wide datasets from ancient human remains, the team was able to identify specific genes that changed during and after the transition from hunting and gathering to farming.
Many of the variants occurred on or near genes that have been associated with height, the ability to digest lactose in adulthood, fatty acid metabolism, vitamin D levels, light skin pigmentation and blue eye color. Two variants appear on genes that have been linked to higher risk of celiac disease but that may have been important in adapting to an early agricultural diet.
Other variants were located on immune-associated genes, which made sense because “the Neolithic period involved an increase in population density, with people living close to one another and to domesticated animals,” said Wolfgang Haak, one of three senior authors of the study, a research fellow at the University of Adelaide and group leader in molecular anthropology at the Max Planck Institute for the Science of Human History.
“Although that finding did not come fully as a surprise,” he added, “it was great to see the selection happening in ‘real time.'”
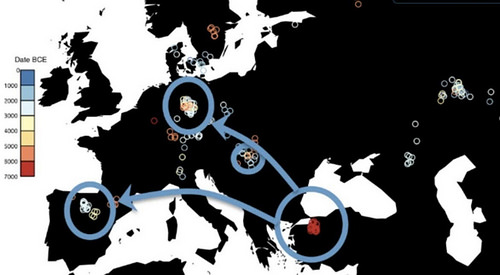
The work also supports the idea that Europe’s first farmers came from ancient Anatolia, in what is now Turkey, and fills in more details about how ancient groups mixed and migrated.
“It’s a great mystery how present-day populations got to be the way we are today, both in terms of how our ancestors moved around and intermingled and how populations developed the adaptations that help us survive a bit better in the different environments in which we live,” said co-senior author David Reich, professor of genetics at HMS. “Now that ancient DNA is available at the genome-wide scale and in large sample sizes, we have an extraordinary new instrument for studying these questions.”
“From an archaeological perspective, it’s quite amazing,” said co-senior author Ron Pinhasi, associate professor of archaeology at University College Dublin. “The Neolithic revolution is perhaps the most important transition in human prehistory. We now have proof that people did actually go from Anatolia into Europe and brought farming with them. For more than 40 years, people thought it was impossible to answer that question.”
“Second,” he continued, “we now have evidence that genetic selection occurred along with the changes in lifestyle and demography, and that selection continued to happen following the transition.”
Prying more from the past
Members of the current team and others have used ancient DNA in the past few years to learn about Neanderthals and the genes they passed to humans, identify ancestors of present-day Europeans, trace migrations into the Americas and probe the roots of Indo-European languages. Studying natural selection, however, remained out of reach because it required more ancient genomes than were available.
“In the past year, we’ve had a super-exponential rise in the number of ancient samples we can study on a genome scale,” said Reich, who is also an associate member of the Broad Institute of Harvard and MIT and a Howard Hughes Medical Investigator. “In September 2014, we had 10 individuals. In this study, we have 230.”
The DNA came from the remains of people who lived between 3,000 and 8,500 years ago at different sites across what is now Europe, Siberia and Turkey. That time span provided snapshots of genetic variation before, during and after the agricultural revolution in Europe.
Among the 230 ancient individuals were 83 who hadn’t been sequenced before, including the first 26 to be gathered from the eastern Mediterranean, where warm conditions usually cause DNA to degrade.
Members of the team used several technological advances to obtain and analyze the new genetic material. For example, they exploited a method pioneered by Pinhasi’s laboratory to extract DNA from a remarkably rich source: a portion of the dense, pyramid-shaped petrous bone that houses the internal auditory organs. In some cases, the bone yielded 700 times more human DNA than could be obtained from other bones, including teeth.
“That changed everything,” said Pinhasi. “Higher-quality DNA meant we could analyze many more positions on the genome, perform more complex tests and simulations, and start systematically studying allele frequency across populations.”
What made the cut
Although the authors caution that sample size remains the biggest limitation of the study, comparing the ancient genomes to one another and to those of present-day people of European ancestry revealed 12 positions on the genome where natural selection related to the introduction of farming in northern latitudes appears to have happened.
“Some of those specific traits have been studied before,” said Reich. “This work with ancient DNA enriches our understanding of those traits and when they appeared.”
Besides the adaptations that appear to be related to diet, pigmentation, immunity and height, the possible selective pressure on other variants was less clear.
“We can guess by looking at the function of the gene, but our power is limited,” said Mathieson. “It’s quite frustrating.”
It’s too early to tell whether some of the variants were themselves selected for or whether they hitched a ride with a nearby beneficial gene. The question pertains especially to variants that seem to be disadvantageous, like increased disease risk.
Being able to look at numerous positions across the genome also allowed the team to examine complex traits for the first time in ancient DNA.
“We can see the evolution of height across time,” said Mathieson.
Researchers had noticed that people from southern Europe tend to be shorter than those from northern Europe. The new study suggests that the height differential arises both from people in the north having more ancestry from Eurasian steppe populations, who seem to have been taller, and people in the south having more ancestry from Neolithic and Chalcolithic groups from the Iberian peninsula, who seem to have been shorter.
The team wasn’t able to draw conclusions about the other complex traits it investigated: body mass index, waist-hip ratio, type 2 diabetes, inflammatory bowel disease and lipid levels.
Reich, for one, hopes researchers will one day have thousands of ancient genomes to analyze. He would also like to see this type of study applied to non-European populations and even to other species.
“It will be interesting to study selection in domesticated animals and to see if there is coevolution between them and the people who were domesticating them,” said Mathieson.
________________________________________
The genomic results confirm that agriculture spread to Europe from Anatolia (ancient Turkey) by human migration or dispersal. Credit: Harvard Medical School
________________________________________________________
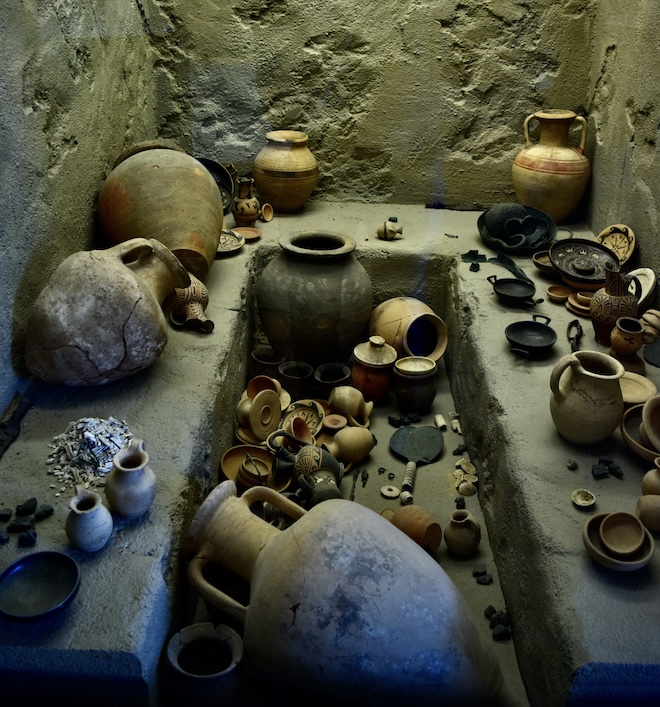
Ancient bone specimen used to obtain samples for genetic testing. Credit: Harvard Medical School
_______________________________________________________
Human Adaptation through Ancient Infections
STANFORD UNIVERSITY MEDICAL CENTER—In another study, genetic material from ancient viral infections were found to be critical to human development, according to researchers at the Stanford University School of Medicine.
They’ve identified several noncoding RNA molecules of viral origins that are necessary for a fertilized human egg to acquire the ability in early development to become all the cells and tissues of the body. Blocking the production of this RNA molecule stops development in its tracks, they found.
The discovery comes on the heels of a Stanford study earlier this year showing that early human embryos are packed full of what appear to be viral particles arising from similar left-behind genetic material.
“We’re starting to accumulate evidence that these viral sequences, which originally may have threatened the survival of our species, were co-opted by our genomes for their own benefit,” said Vittorio Sebastiano, PhD, an assistant professor of obstetrics and gynecology. “In this manner, they may even have contributed species-specific characteristics and fundamental cell processes, even in humans.”
Sebastiano is a co-lead and co-senior author of the study, which will be published online Nov. 23 in Nature Genetics. Postdoctoral scholar Jens Durruthy-Durruthy, PhD, is the other lead author. The other senior author of the paper is Renee Reijo Pera, PhD, a former professor of obstetrics and gynecology at Stanford who is now on the faculty of Montana State University.
Sebastiano and his colleagues were interested in learning how cells become pluripotent, or able to become any tissue in the body. A human egg becomes pluripotent after fertilization, for example. And scientists have learned how to induce other, fully developed human cells to become pluripotent by exposing them to proteins known to be present in the very early human embryo. But the nitty-gritty molecular details of this transformative process are not well understood in either case.
An ancient infection
The researchers knew that a type of RNA molecules called long-intergenic noncoding, or lincRNAs, have been implicated in many important biological processes, including the acquisition of pluripotency. These molecules are made from DNA in the genome, but they don’t go on to make proteins. Instead they function as RNA molecules to affect the expression of other genes.
Sebastiano and Durruthy-Durruthy used recently developed RNA sequencing techniques to examine which lincRNAs are highly expressed in human embryonic stem cells. Previously, this type of analysis was stymied by the fact that many of the molecules contain highly similar, very repetitive regions that are difficult to sequence accurately.
They identified more than 2,000 previously unknown RNA sequences, and found that 146 are specifically expressed in embryonic stem cells. They homed in on the 23 most highly expressed sequences, which they termed HPAT1-23, for further study. Thirteen of these, they found, were made up almost entirely of genetic material left behind after an eons-ago infection by a virus called HERV-H.
HERV-H is what’s known as a retrovirus. These viruses spread by inserting their genetic material into the genome of an infected cell. In this way, the virus can use the cell’s protein-making machinery to generate viral proteins for assembly into a new viral particle. That particle then goes on to infect other cells. If the infected cell is a sperm or an egg, the retroviral sequence can also be passed to future generations.
HIV is one common retrovirus that currently causes disease in humans. But our genomes are also littered with sequences left behind from long-ago retroviral infections. Unlike HIV, which can go on to infect new cells, these retroviral sequences are thought to be relatively inert; millions of years of evolution and accumulated mutations mean that few maintain the capacity to give instructions for functional proteins.
After identifying HPAT1-23 in embryonic stem cells, Sebastiano and his colleagues studied their expression in human blastocysts — the hollow clump of cells that arises from the egg in the first days after fertilization. They found that HPAT2, HPAT3 and HPAT5 were expressed only in the inner cell mass of the blastocyst, which becomes the developing fetus. Blocking their expression in one cell of a two-celled embryo stopped the affected cell from contributing to the embryo’s inner cell mass. Further studies showed that the expression of the three genes is also required for efficient reprogramming of adult cells into induced pluripotent stem cells.
Sequences found only in primates
“This is the first time that these virally derived RNA molecules have been shown to be directly involved with and necessary for vital steps of human development,” Sebastiano said. “What’s really interesting is that these sequences are found only in primates, raising the possibility that their function may have contributed to unique characteristics that distinguish humans from other animals.”
The researchers are continuing their studies of all the HPAT molecules. They’ve learned that HPAT-5 specifically affects pluripotency by interacting with and sequestering members of another family of RNAs involved in pluripotency called let-7.
“Previously retroviral elements were considered to be a class that all functioned in basically the same way,” said Durruthy-Durruthy. “Now we’re learning that they function as individual elements with very specific and important roles in our cells. It’s fascinating to imagine how, during the course of evolution, primates began to recycle these viral leftovers into something that’s beneficial and necessary to our development.”
__________________________________________
The first study was supported by U.S. National Science Foundation HOMINID grant BCS-1032255, U.S. National Institutes of Health grant GM100233, European Research Council (ERC) starting grants ADNABIOARC (263441) and APGREID, an Irish Research Council ERC support grant, the Irish Research Council for Humanities and Social Sciences, the Australian Research Council (DP130102158), the German Research Foundation (DFG) grants KR 4015/1-1 and AL 287/14-1, the Human Frontier Science Program (LT001095/2014-L), the Russian Foundation for Basic Research (5-06-01916), the Russian Foundation for the Humanities (15-11-63008), and a state grant from the Ministry of Education and Science of the Russian Federation (#33.1195.2014/k).
In the second study, other Stanford authors are postdoctoral scholars Mark Wossidlo, PhD, Jonathan Davila, PhD, and Moritz Mall, PhD; research associate Diana Cepeda, PhD; former postdoctoral scholar Jun Cui, PhD; graduate student Edward Grow; Wing Wong, PhD, professor of statistics and health research; and Joanna Wysocka, PhD, professor of chemical and systems biology and of developmental biology.
That study was supported by the National Institutes of Health (grants 1U54HD068158 and 1U01HL100397) and the California Institute for Regenerative Medicine.
Stanford’s Department of Obstetrics and Gynecology also supported the work.
_____________________________________________________
In addition, the latest Popular Archaeology ebook is now available.
______________________________________________
Travel and learn with Far Horizons.
____________________________________________
This richly illustrated issue includes the following stories: Two remarkable discoveries that are shedding light on human beginnings in Africa; a traveling exhibit and an archaeological site that show how knowledge is more valuable than gold; a Spanish cave and a unique burial that are offering a tantalizing glimpse on the lives of Ice Age hunter-gatherers in Europe; the stunning visual reconstruction of an ancient Roman town; enlightening new finds at a remarkably well-preserved site of ancient Hellenistic-Roman culture overlooking the Sea of Galilee; rare finds that are shedding light on occult practices among ancient Greeks in Sicily; and an overview of the overwhelmingly rich archaeological heritage of Britain. Find it on Amazon.com.